Six month was not quite the amount of time I anticipated between the third and fourth edition, but I finally managed to upload edition 1.4.7-0 of my Groovy Cheminformatics book. The first three editions sold 37 copies, including two for myself.

The Oscar4 paper (CC-BY, just like the screenshots of the paper below) was out already some days now, but the formatting has finished: I spotted a rogue http:// in the code example b) in Appendix B: I’ll see what I can do about that, but the API might evolve a bit anyway.
I reported earlier how to I uploaded the ChemPedia (RIP) data onto Kasabi. But for ChEMBL-RDF I have used the pytassium tool, not just because it has a cool name :) I discovered yesterday, however, that I did not write down in this lab notebook, what steps I needed to take to reproduce it. And I just wanted to uploaded new triples to the ChEMBL-RDF data set on Kasabi.
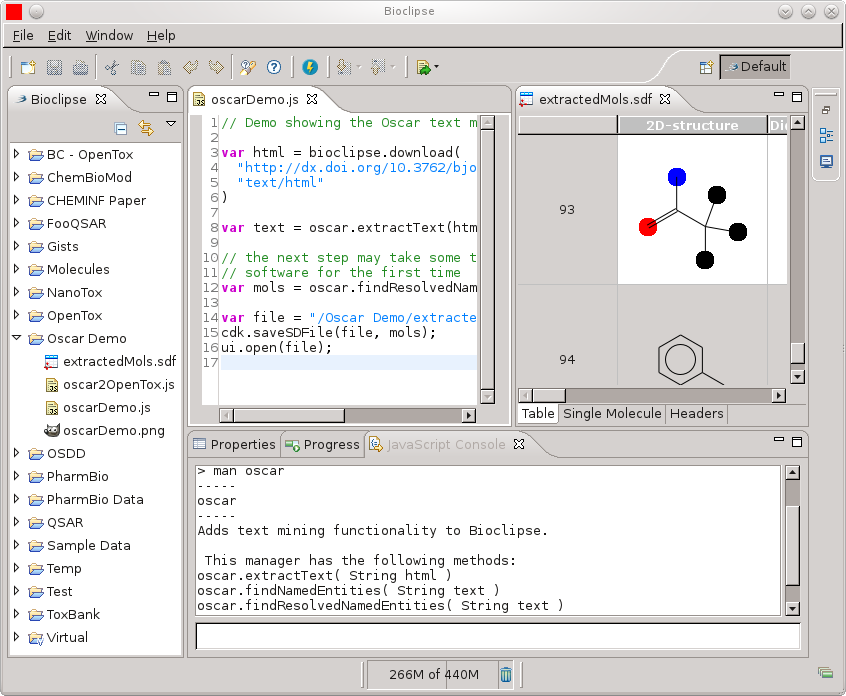
Almost a year ago I started a position with Peter Murray-Rust to work on Oscar for three months (see this overview of results; a paper by the full Oscar team (Sam, David, Dan, Lezan) is pending, and I’m really happy to have been able to contribute bits to the project). Since then, I have had little time :( That’s how it goes, with post-hopping, unfortunately. One thing I did do after that, was write a Bioclipse plugin.
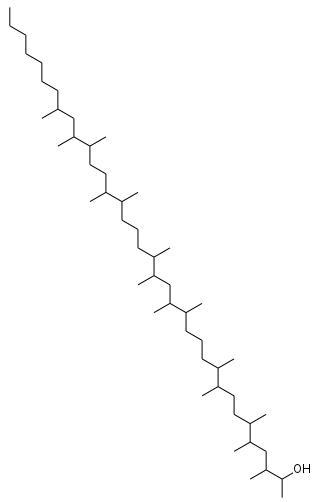
About two weeks ago, the ChemConnector blog reported an InChIKey collosion detected by Prof. Goodman . Unlike the previous collision, this one was based solely on the graph and not on stereochemistry. The two molecules both have the InChIKey OCPAUTFLLNMYSX-UHFFFAOYSA-N: The compounds are really different, the molecular formulas are C 50 H 102 O and C 57 H 114 O respectively.
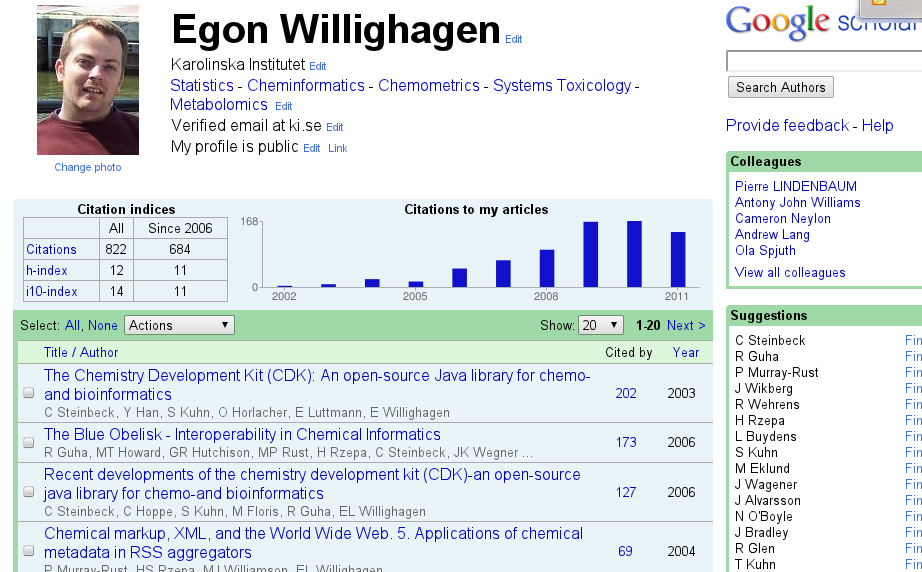
Web of Science is my de facto standard for citation statistics (I need these for VR grant applications), and defines the lower limit of citations (it is pretty clean, but I do have to ping them now and then to fix something). The public front-end of it is Researcher ID. There is an Microsoft initiative, which looks clean but doesn’t work on Linux for the nicer things, but the coverage of journals is pretty bad in my field, giving a biased
Update: the fourth edition is out.
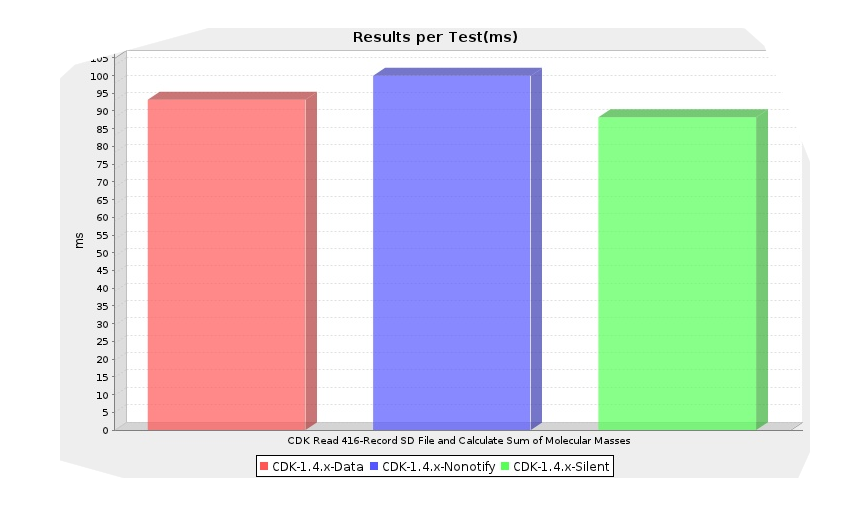
I cannot find the bug report just now, but the CDK has an open problem with change even notification, where the nonotify classes still caused change event to be sent around. This was because the nonotify classes extended in a wrong way the data classes. So, I worked today on copying the data class implementations into a new implementation, not extending the data classes, while removing the listener code: the silent module.

Kasabi is a new, RDF hosting service by Talis. It’s still in beta, and I have been testing their beta service with the RDF version I created of ChemPedia Substances (the now no longer existing cool web service from MetaMolecular to draw and name organic molecules). Kasabi makes the RDF data available via a few APIs, depending on the APIs selected by the uploader. I picked all five of them, just to see how things work.
Update 2021-02 : this post is still the second-most read post in my blog. Welcome! Some updates: Ammar Ammar in our BiGCaT group has set up a new SPARQL endpoint. Please use and tweet. blog, or otherwise let others now how you use the ChEMBL RDF. Since this post I have blogged a lot more about ChEMBL. Update : this work is now written down in this paper.

Update: the fourth edition is out.
