Notes to self on web map-style tree viewers. The basic idea is to use Google Maps or Leaflet to display a tree. Hence we need to compute tiles. One approach is to use a database that supports spatial queries to store the x,y coordinates of the tree. When we draw a tile we compute the coordinates of that tile, based on position and zoom level, do a spatial query to extract all lines that intersect with the rectangle for that tile, and draw those.
iPhylo

Continuing my on-again off-again relationship with the Semantic Web, I stumbled across a cool approach to visualising the results of SPARQL queries.

TL;DR By using bookmarklets and a central annotation store, we can build a system to annotate any biodiversity database, and display those annotations on those databases. A couple of weeks ago I was at GBIF meeting in Copenhagen, and there was a discussion about adding a new feature to the GBIF portal. The conversation went something like this: Resources are limited, and adding new features to a project can be difficult.

Say what you will about Elsevier, they are certainly exploring ways to re-imagine the scientific article. In a comment on an earlier post Fabian Schreiber pointed out that Elsevier have released an app to display phylogenies in articles they publish. The app is based on jsPhyloSVGand is described here.
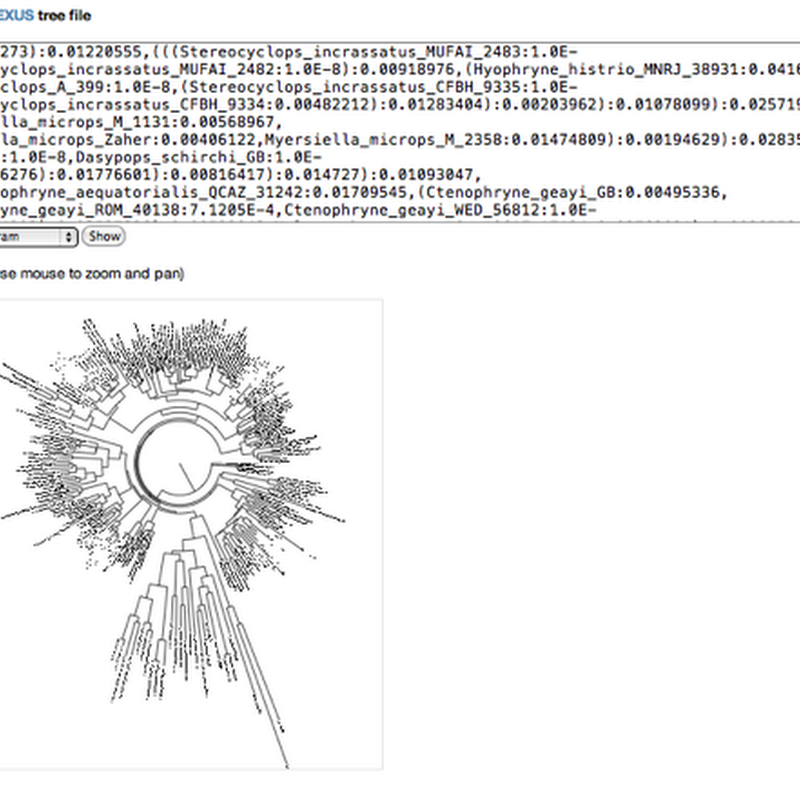
Following on from the SVG experiments I've started to put some of the Javascript code for displaying phylogenies on Github. Not a repository yet, but as gists, little snippets of code. Mike Bostock has created http://bl.ocks.org/ which makes it possible to host gists as working examples, so you can play with the code "live". The first gist takes a Newick tree, parses it and displays a tree.
One visualisation method I keep coming back too is the treemap. Each time I experiment with them I learn a little bit more, but I usually end up abandoning them (with the exception of using quantum treemaps to display bibliographic data). But they keep calling me back.
One thing I'm increasingly conscious of is that I've a lot of demos and toy projects hanging around and the code for most of these isn't readily available. So, I plan to clean these up and put them in GitHub so others can explore the code, and reuse it if they see fit. First up is the code to create a HTML+Javascript clone of Nature's iPhone app, as described in an earlier post. There's a live version of the clone here here.

The bulk of the Biodiversity Heritage Library's content is available as DjVu files, which package together scanned page images and OCR text. Websites such as BHL or my own BioStor display page images, but there's no way to interact with the page content itself.

One of the more glaring limitations of my BHL viewer described in the previous post is that it can take a while to load all the page thumbnails (there can be hundreds). Given that one of the original motivations for this project was a faster viewer, this kinda sucks. What I'd like to do is load the thumbnails only when I need them, rather than all at once at the start -- in other words I'd like to implement lazy loading.

I've just discovered Nicolas Garcia Belmonte's JavaScript Information Visualization Toolkit (JIT). Wow! This is very cool stuff (and no Flash). To quote from the web site: Nicolas also links to a talk by Tamara Munzner, which I've embedded below to remind myself to watch it.