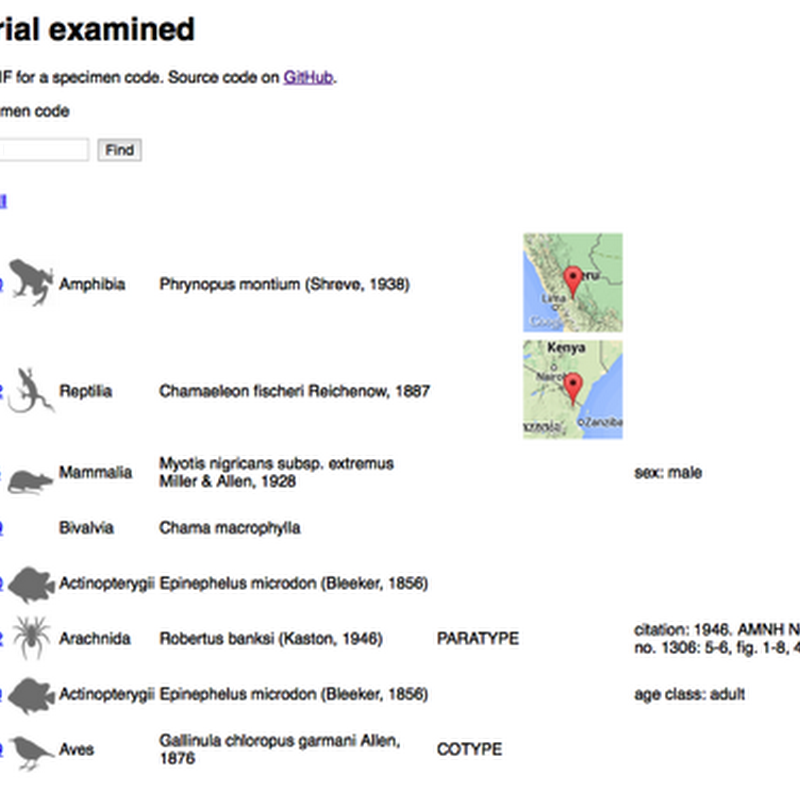
I've put together a working demo of some code I've been working on to discover GBIF records that correspond to museum specimen codes. The live demo is at http://bionames.org/~rpage/material-examined/ and code is on GitHub. To use the demo, simply paste in a specimen code (e.g., "MCZ 24351") and click Find and it will do it's best to parse the code, then go off to GBIF and see what it can find.