
I just returned from a week in Barcelona where I attended the Nextflow Summit and nf-core hackathon, and I can hardly contain my excitement for the near term future of bioinformatics, computational biology, and open science in general.

I just returned from a week in Barcelona where I attended the Nextflow Summit and nf-core hackathon, and I can hardly contain my excitement for the near term future of bioinformatics, computational biology, and open science in general.
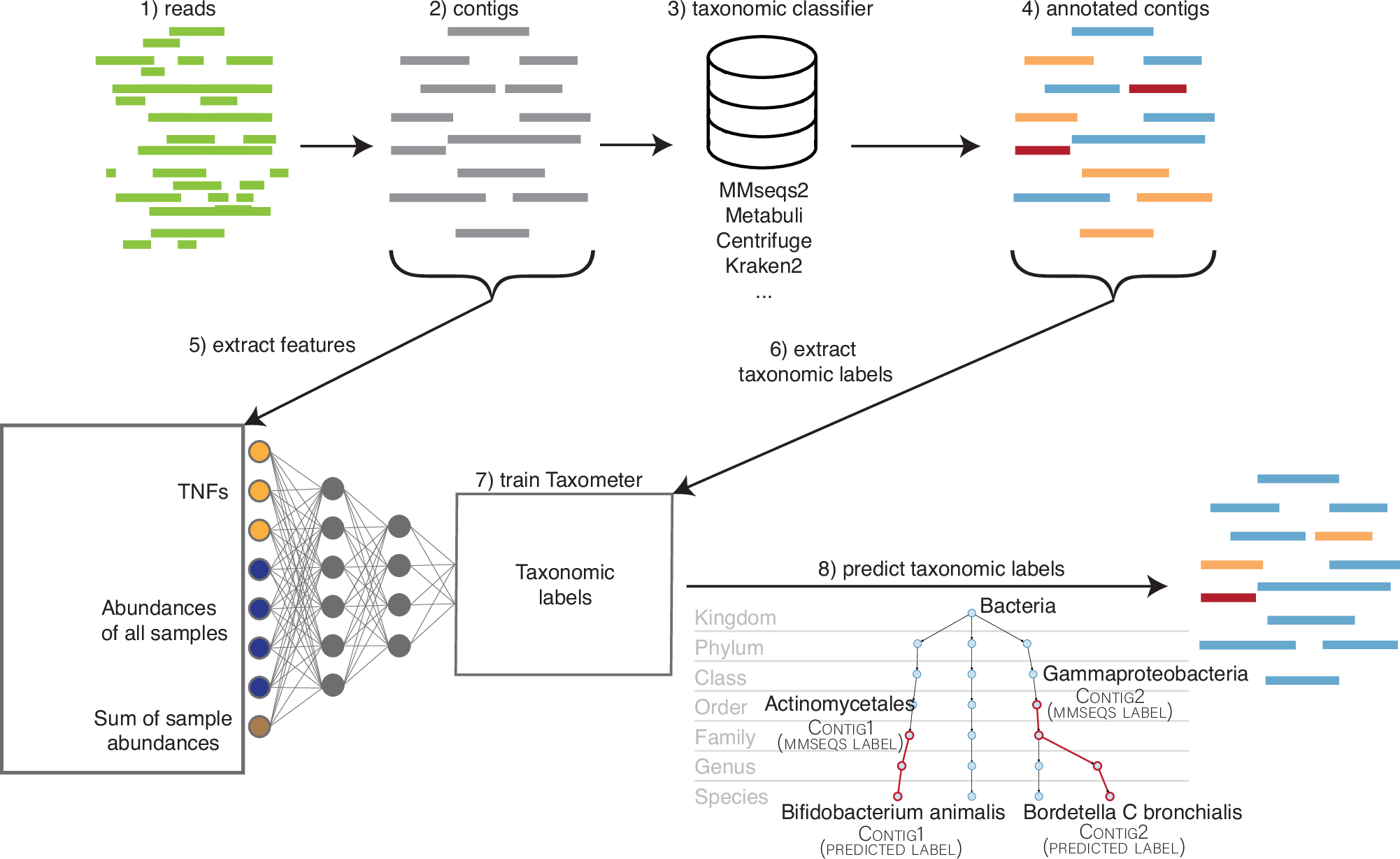
This week’s recap highlights protein design with RoseTTAFold, surveillance with wastewater sequencing, T2T human genomes, Vitessce for visualization of multimodal spatial single-cell data, and Taxometer for taxonomic classification of metagenomics contigs.

This week’s recap highlights a new Nextflow workflow for calculating polygenic scores with adjustments for genetic ancestry, a paper demonstrating that whole exome plus imputation on more samples is more powerful than whole genome sequencing for finding more trait associated variants, a new deep-learning-based splice site predictor that improves spliced alignments, a new method for accurate community profiling of large metagenomic datasets, and

I am in the middle of writing a review / perspectives paper. One that I’m confident will be exciting once we get it published. Some sections of the review cover subject matter at the outer periphery of my expertise.
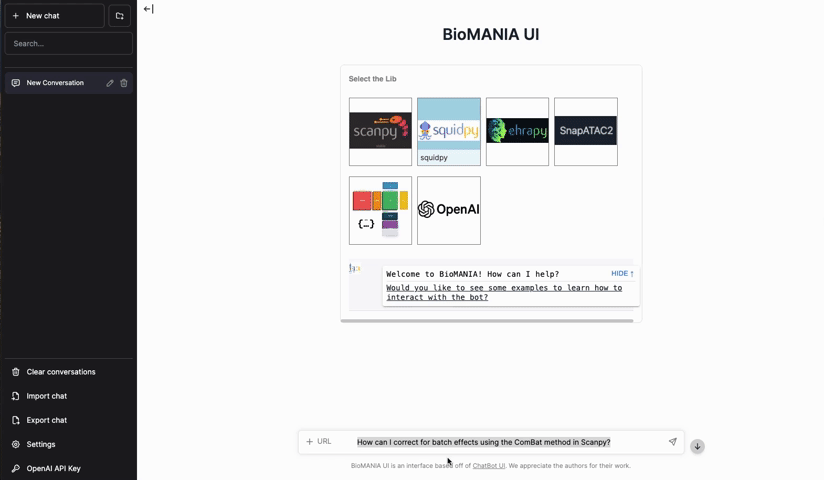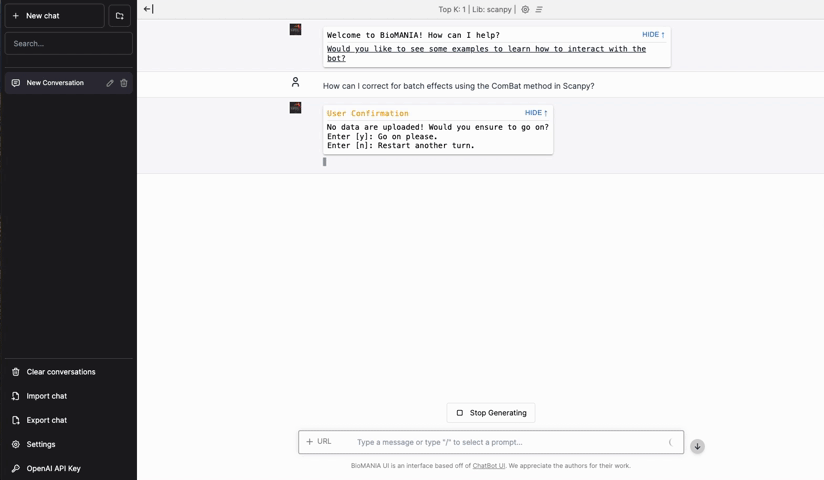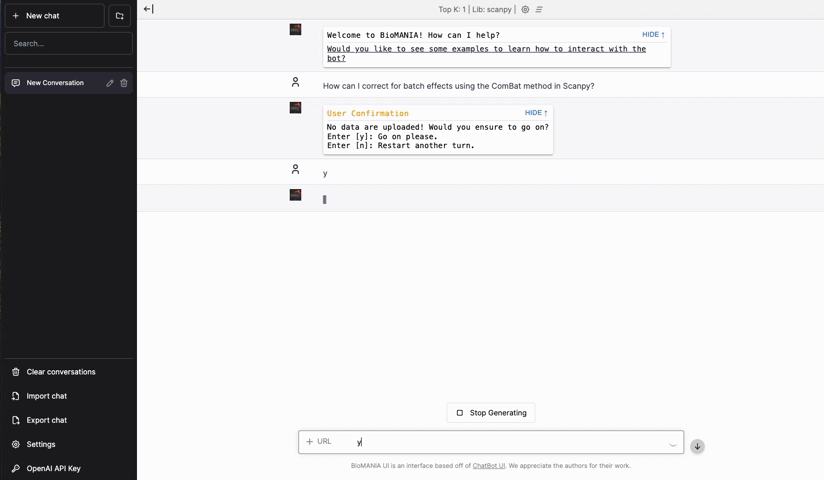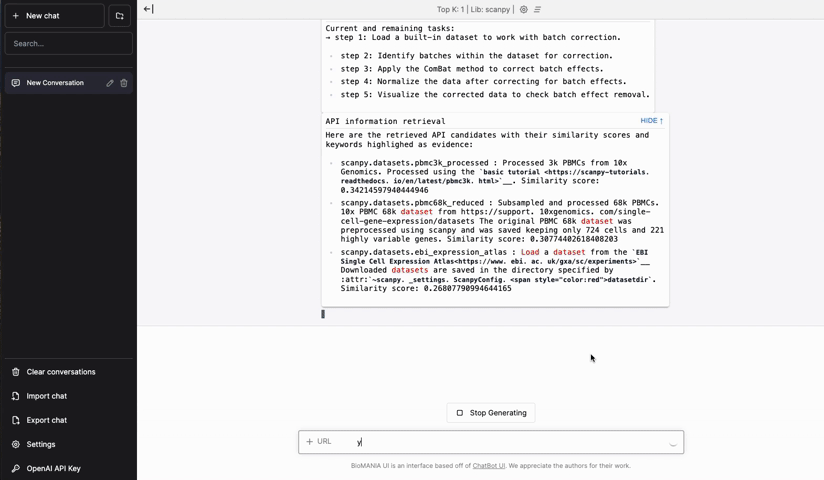
This week’s recap highlights a new method for gene-level alignment of single-cell trajectories, an R package for integrating gene and protein identifiers across biological sequence databases, characterization of SVs across humans and apes, universal prediction of cellular phenotypes, a method to quantify cell state heritability versus plasticity and infer cell state transition with single cell data, and a new AI-driven, natural language-oriented

This week’s recap highlights a new multispecies codon optimization method, personalized pangenome references with vg, a commentary on the wild west of spike-in normalization, a new pipeline for comprehensive and scalable polygenic scoring across ancestrally diverse populations, a paper showing deep learning / transformer-based methods don’t outperform simple linear models for predicting gene expression after genetic perturbations, and finally, a
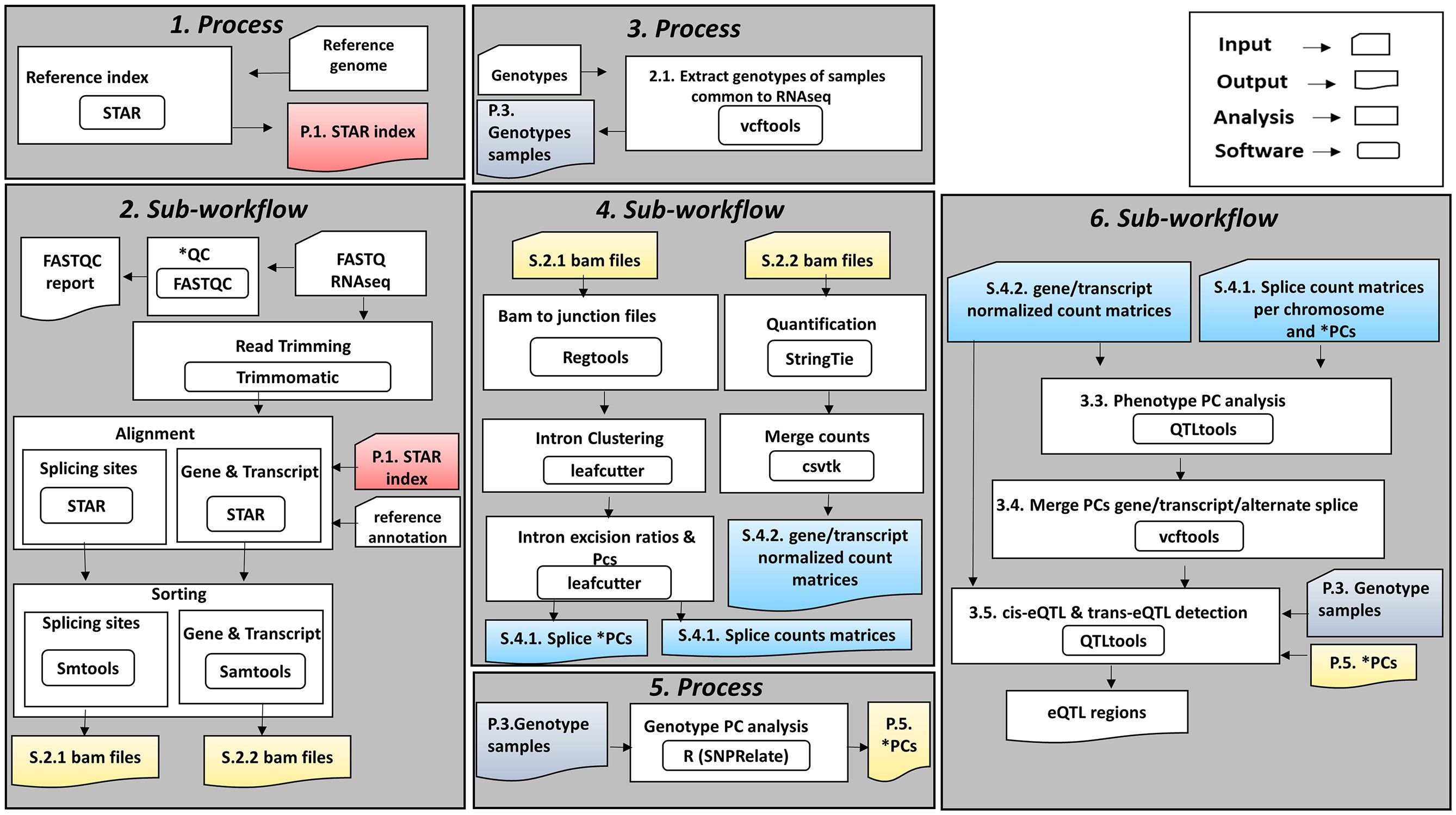
This week’s recap highlights a Nextflow pipeline for eQTL detection, an end-to-end pipeline for spatial transcriptomics (visium) data analysis, a method for identification of perturbed cell types in single cell RNA-seq data, a method for guide assignment in single-cell CRISPR screens, a tool for on-target/off-target analysis of gene editing outcomes, and “digital microbes” for collaborative team science on emerging microbes.
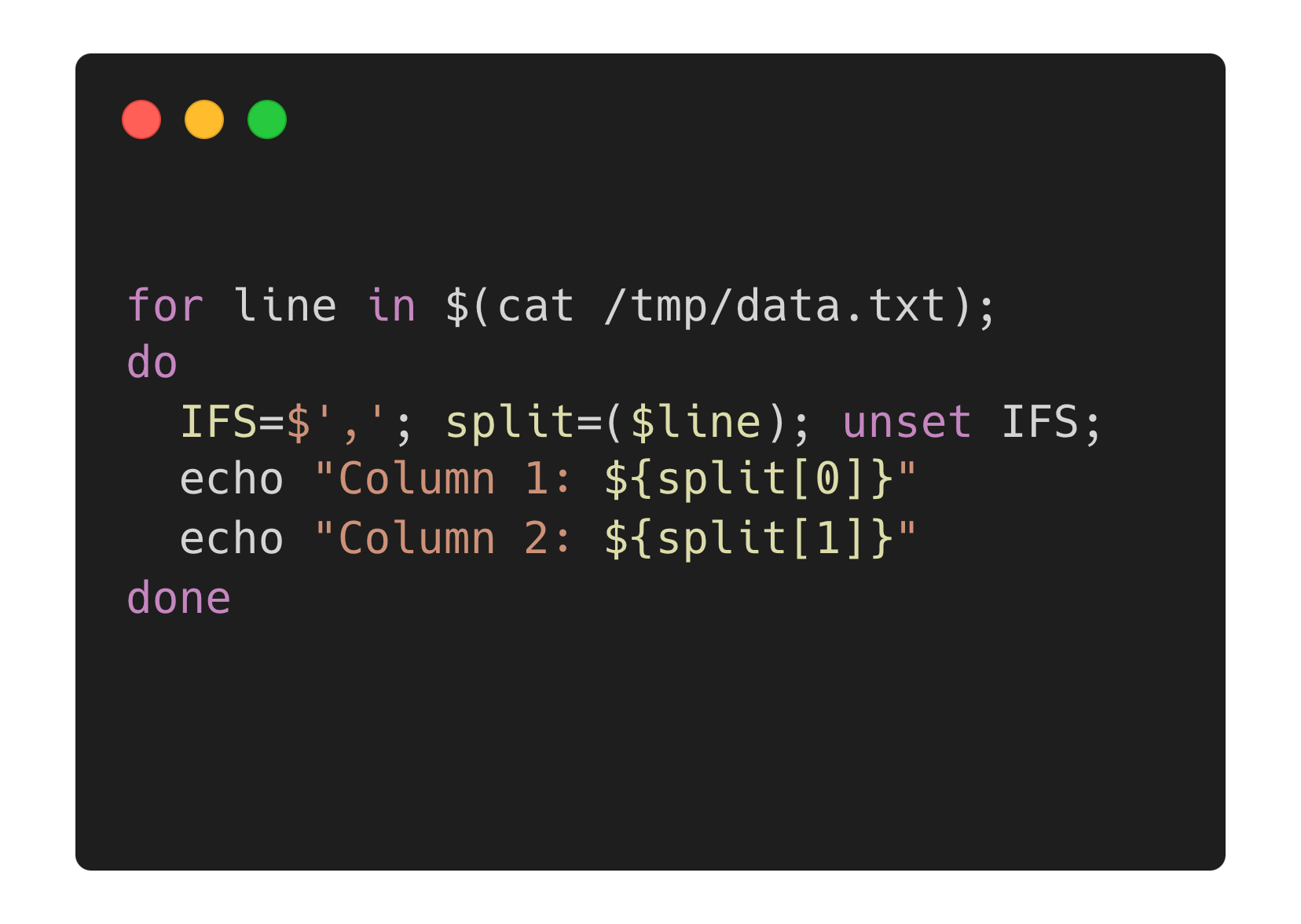
I wrote my first public blog post in 2009. I started Getting Genetics Done to share what I was learning at the end of my PhD/postdoc through my first few years as faculty. Some of the earliest posts were simple, such as how to write and run a simple Perl script, to bigger topics like why it’s usually a bad idea to categorize continuous variables in a linear model.

I recently stumbled across Phil Ewels’s ~18 minute nf-core/bytesize talk on Excalidraw: For years I’ve been using draw.io for making flowcharts and diagrams for documentation, papers, presentations, and for general brainstorming and communication with my team, clients, and collaborators.1 Excalidraw (excalidraw.com) looks like an attractive alternative.
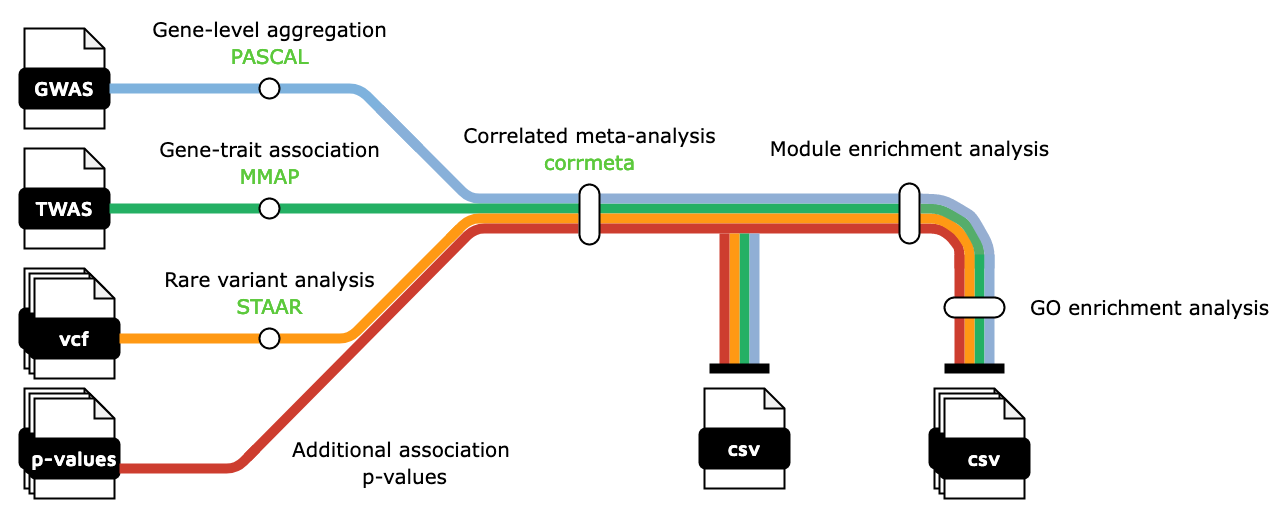
This week’s recap highlights a new nf-core workflow for multi-omics trait association studies, a new tool for linking genotype to phenotype (G2P) by directly sequencing alleles from CRISPR base editing experiments, the SplitsTree app for interactive analysis and visualization using phylogenetic trees and networks, mapping cellular interactions from spatially resolved transcriptomics data, a study of marine microbial diversity and bioprospecting