NSF reorg, Science in 2050, 2025 LLM recap, R+Python, R Data Scientist, virtual cells, genomics in 2026, Claude Code course, AI and labor, how uv got so fast, Anthropic/biotech, papers+preprints

NSF reorg, Science in 2050, 2025 LLM recap, R+Python, R Data Scientist, virtual cells, genomics in 2026, Claude Code course, AI and labor, how uv got so fast, Anthropic/biotech, papers+preprints

Part 3 in a series of posts going back 14 years
On finding joy in slow, imperfect code in an age of copilots, agents, and chatbots. 1.2k words, 5 min reading time.

AI creates an education challenge, not just a job crisis. We haven't built systems to help people to continue learning and connect them to new opportunities.

If an AI produces something useful it's because of your own skill in model choice, prompting, and steering. If not, it's because the model is a useless lying machine that can't follow directions.
R updates (R Data Scientist, R Weekly), AI accelerating wet lab bio research, Docker hardened images, LLMs in review, machinal bypass, science funding, new papers.

The "Machinal Bypass," when AI becomes a shortcut around the work that makes us human, and why some tasks should feel a little hard.
Writing better code without AI, AI in peer review, local LLMs, Biothreat Benchmark Generation, R updates (R Data Scientist, RWeekly, R Works), red-teaming an AI vending machine, new papers

Biotech policy: Recent policy discussions echo NSCEB recommendations across investment, defense, and data.
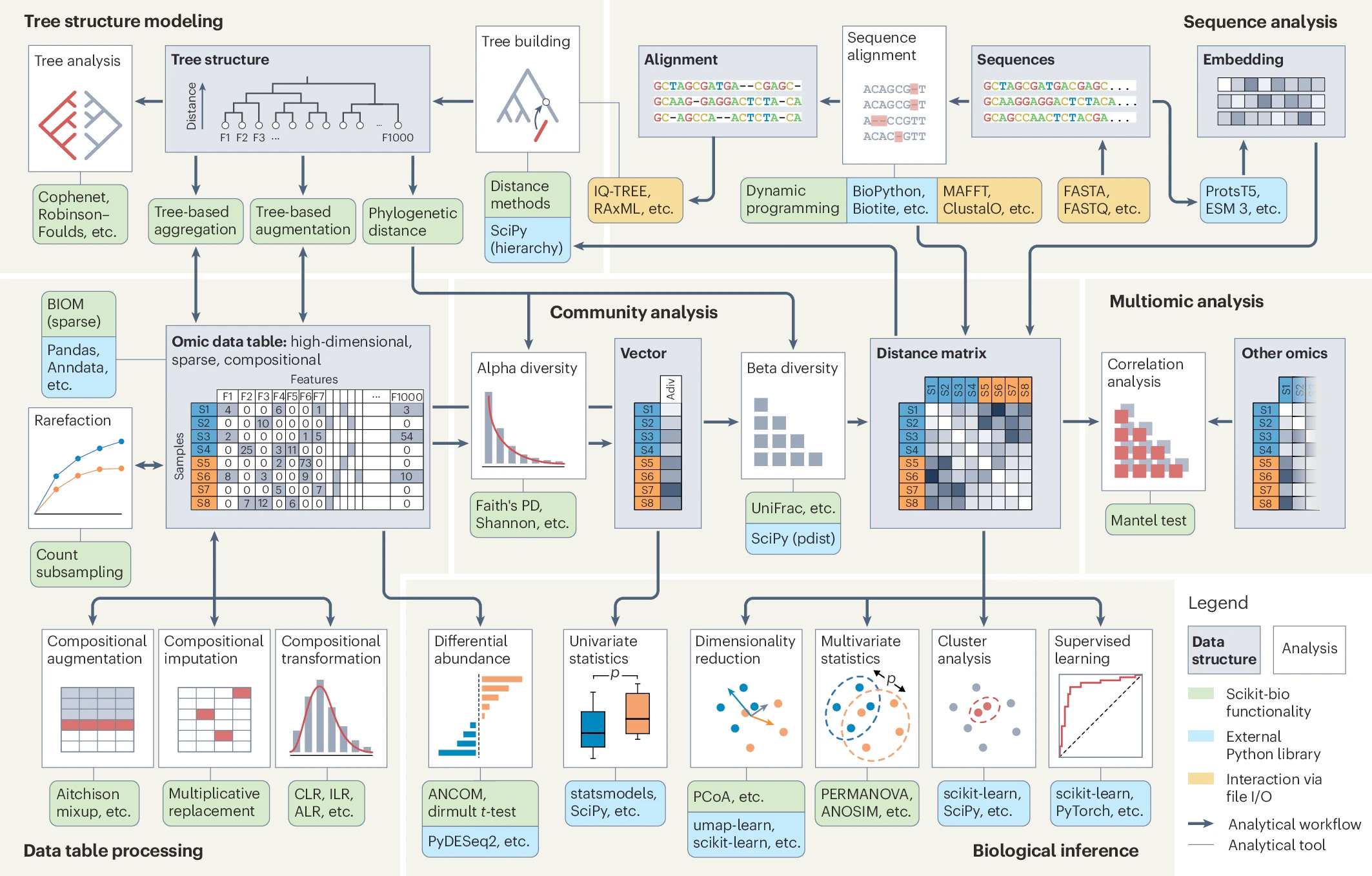
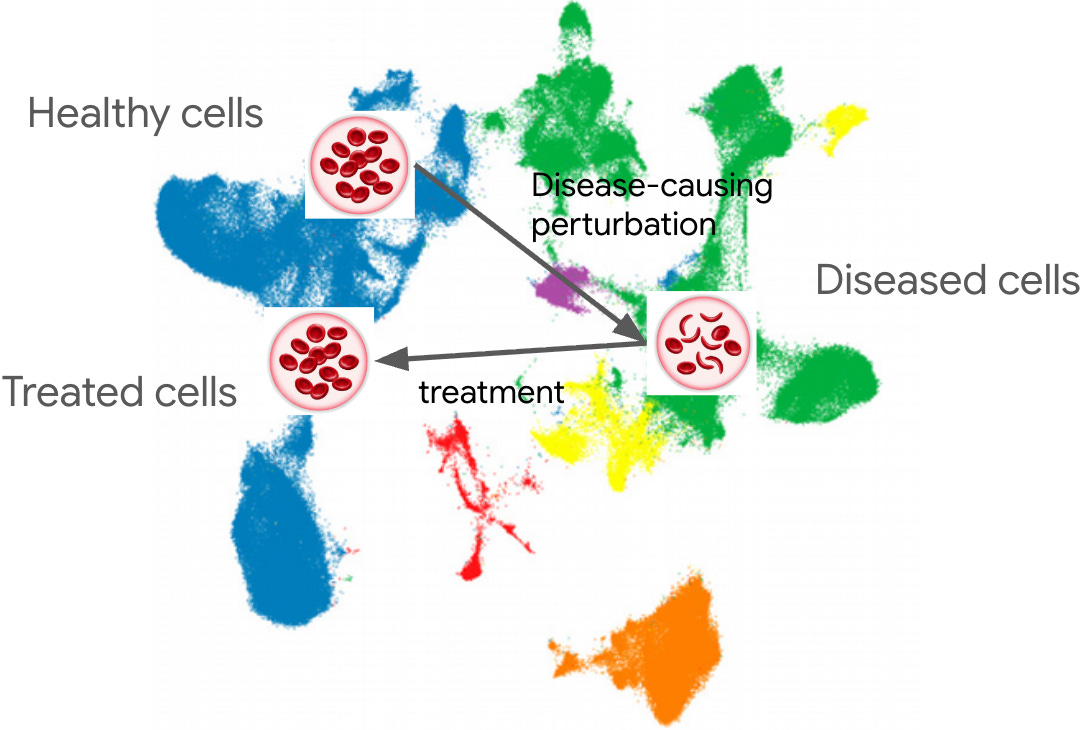
The "missing middle" for omics data analysis in Python
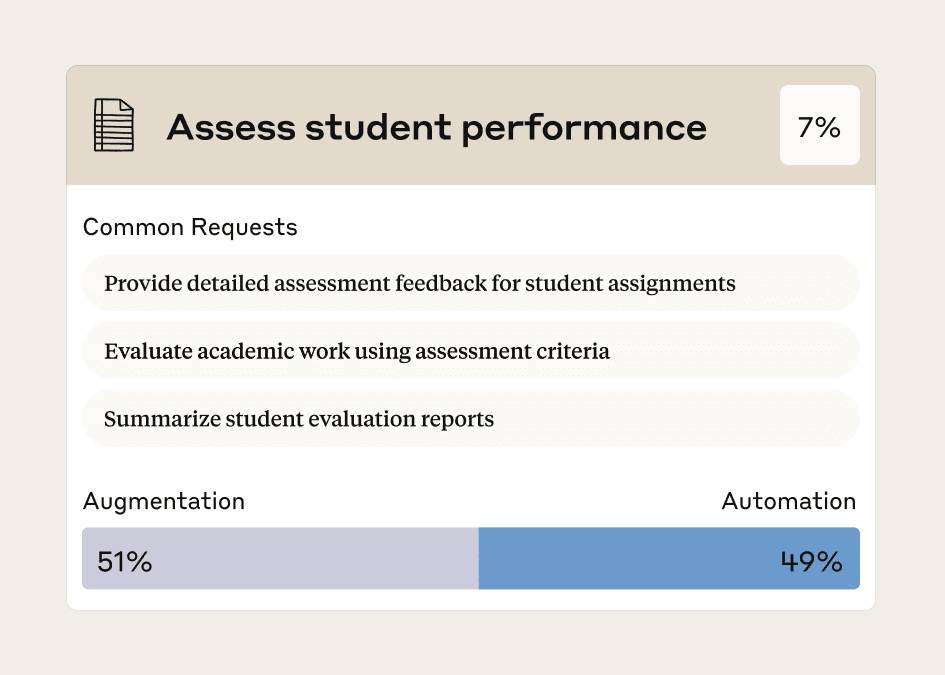

The Anthropic Education Report showed 50% of Claude conversations about grading delegated assessment to the AI. How does one actually design AI-resistant assignments?